Genomic Stability QC
Detecting Genomic Aberrations in iPSC Clones Using Label-free Brightfield Imaging and AI-based AnalysisControlling the genetic integrity of iPSC clones during expansion and maintenance is crucial, as it (1) determines the cells' potential to differentiate into the desired functional target cell types and (2) ensures viability of the resulting cultures. It would be highly desirable to establish a cost-effective and fast method to routinely check for genetic integrity in laboratories using iPSC technology.
In order to explore the viability of such a method, the VAIDR team collaborated with researchers from the iPSC technology platform at the Max-Delbrück-Centrum Berlin, who provided samples for method development and testing. In this application note, we demonstrate the concept of a morphology-based quality control that is able to predict whether a genetic aberration has occurred.

“My group is focused on using the latest technologies to advance the field of stem cell research. We will continue to use AI-based image analysis to support our automated workflows.”
–Sebastian Diecke, Technology Platform Leader at Max-Delbrück-Centrum
Sebastian Diecke leads the pluripotent stem cell group at Max-Delbrück-Centrum Berlin, Germany. His research centers on advanced methodologies, like fully automated cell culture. With this work, he is laying the foundations for a future in which novel cures are based on therapeutic cell products, reliably produced at an industrial scale.
Sebastian's group supported this proof-of-concept study with planning and and execution of the cell culture experiments.
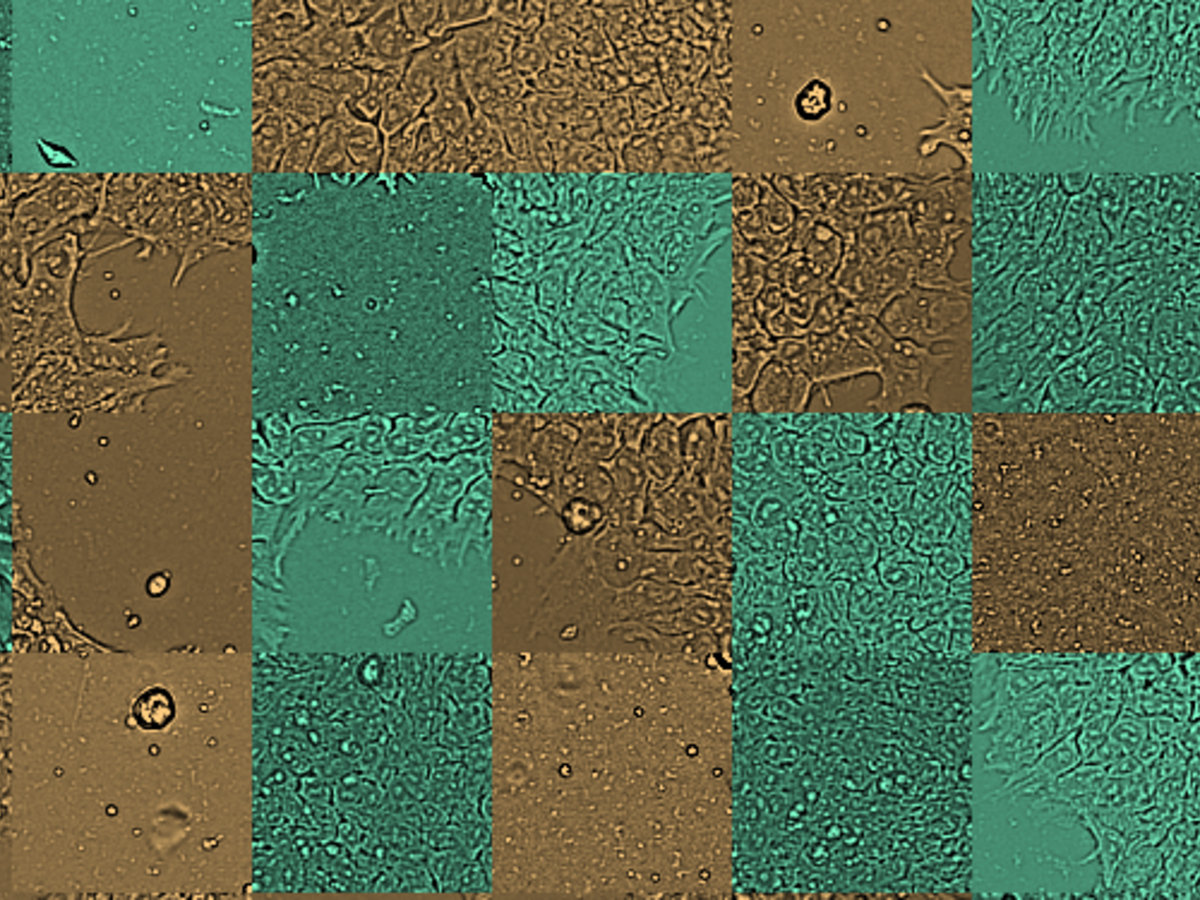
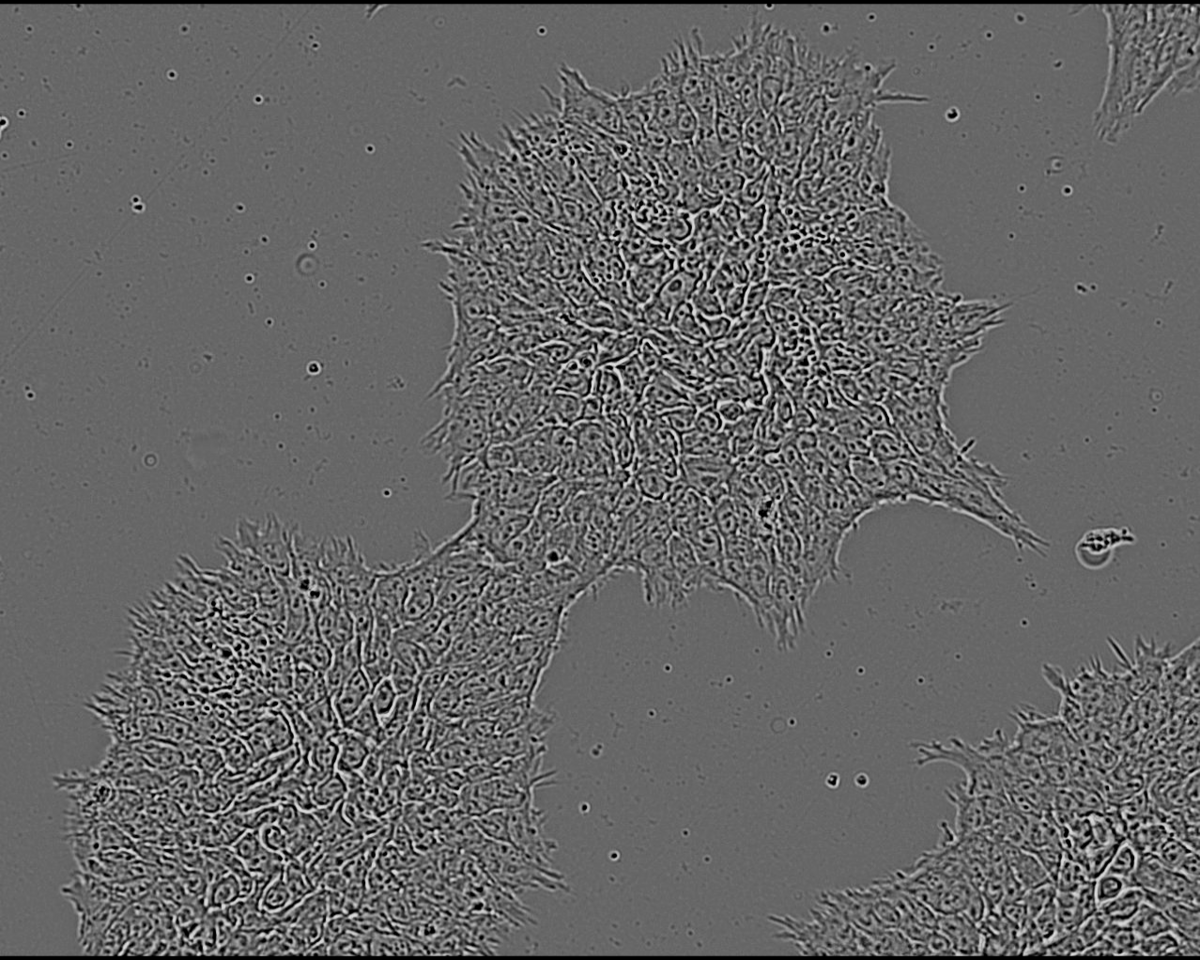
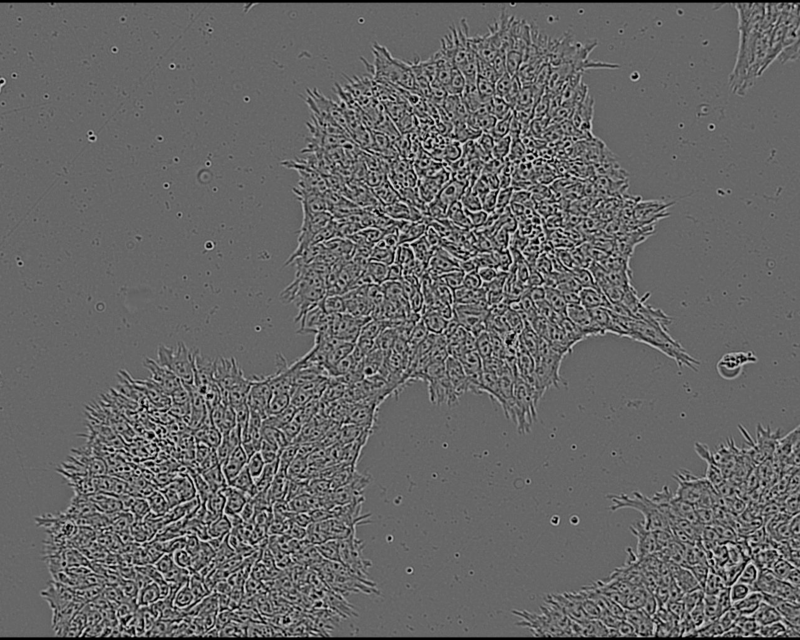
Researchers at the Technology Platform of the Max-Delbrück-Centrum cultured 8 clones from one iPSC line and characterized them for genomic aberrations for reference using SNP Array for Virtual Karyotyping. Three clones (A1, A2, A3) were genetically normal while three others (B1, B2, B3) had an aberration on chromosome 20. The remaining two clones X and Y were also characterized for reference but their genetic status was not revealed to the VAIDR analysis team until after they had made their prediction. The cells were cultured for 24h and then imaged with the VAIDR microscope using only the brightfield channel at 40x magnification. For each clone, several replicate wells were prepared in 6-well plates and each well was imaged at 49 locations.
This study is an example of a classification task: The aim is to develop a method that assigns unknown samples to one of several classes. In the present case, images from routine culture should be classified as showing genetically normal or aberrant cells. A key requirement is that no a priori knowledge about the visual differences between conditions (normal vs aberrant) has to be available. Rather, the method should be constructed only from the images of known genetically normal and aberrant cells. VAIDR offers a dedicated workflow for this kind of scenario.
The VAIDR team used the built-in classification workflow to train a classification method that distinguished genetically normal clones from clones with an aberration on chromosome 20. For the training, only clones A1, A2, A3, B1, B2 and B3 were used. This left the remaining clones X and Y to be used as a completely independent test for challenging the method.
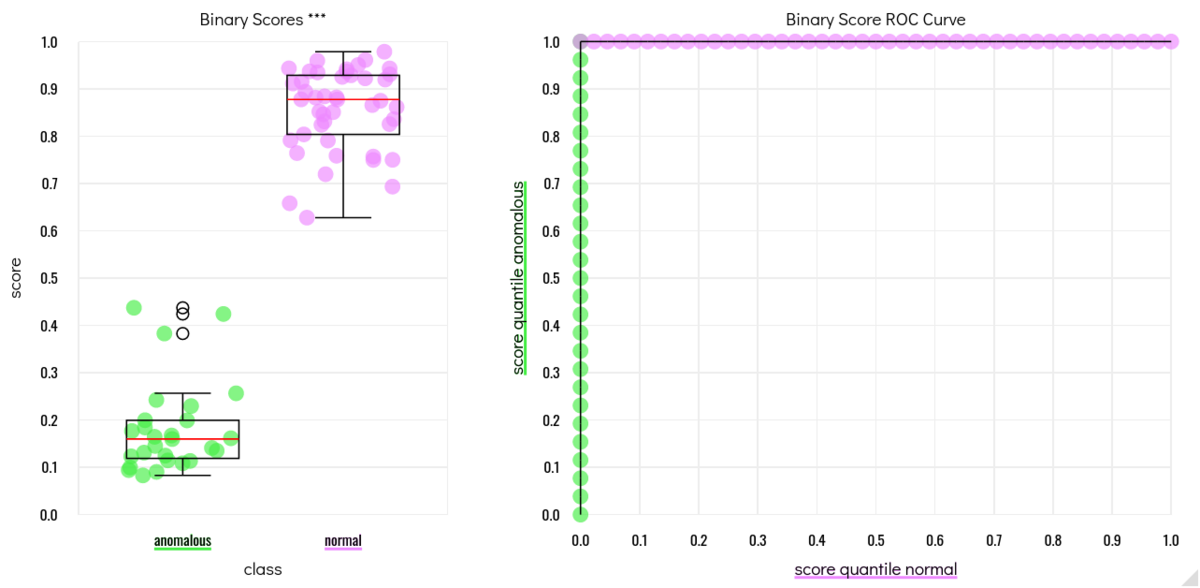
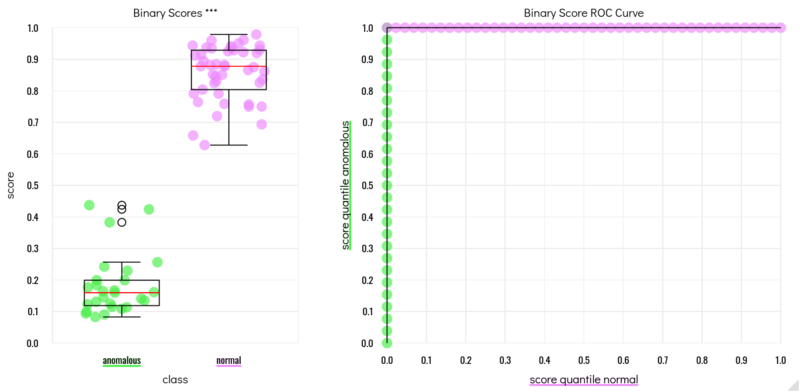
Results of cross-validation during training. For training the AI model, each clone was left out once, and the model was trained on all remaining clones. Then, the left-out clone was scored with the model. The score plots and ROC on the left show, that the training went perfectly: Each clone was correctly mapped as being genetically normal or having an aberration on chromosome 20. After this positive validation result, the final model was trained using all clones at once. This type of cross-validation is performed automatically by the VAIDR classification workflow for each new classification method, and a validation report containing performance statistics and plots is generated automatically.
After successful method development and validation, the VAIDR analysis team applied the method to all clones, in particular to the ones whose genetic status was still blinded to the analysis team at that time point. Each clone received a classification score, where a high score corresponds to the detection of a genetic aberration on chromosome 20 and a low score indicates a genetically normal clone.
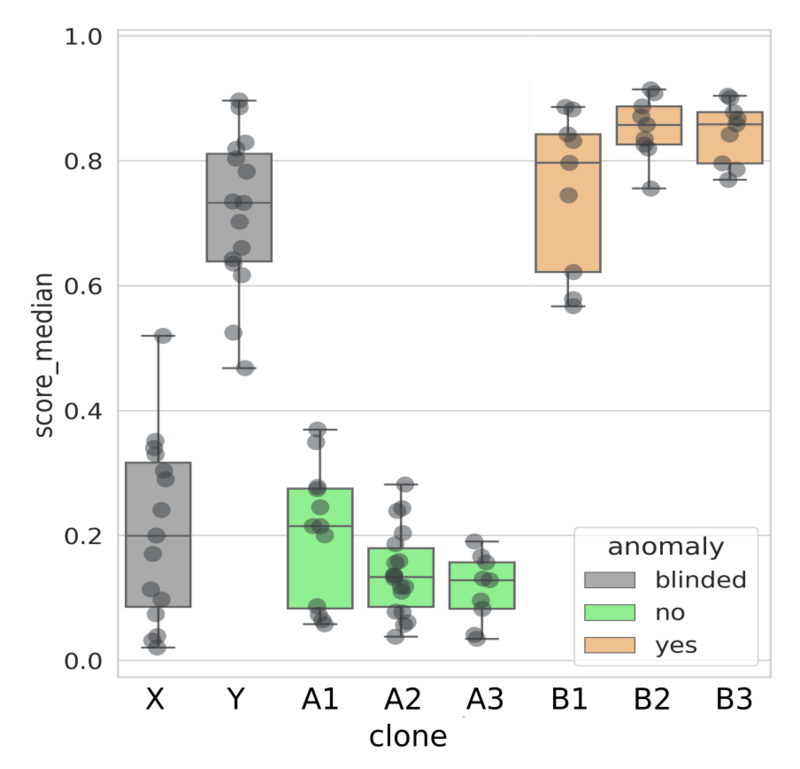
Analysis results of scoring images from all clones on with VAIDR classification method trained on genetically normal clones A1, A2 and A3 vs the clones with an aberration on Chromosome 20 (B1, B2, B3). The clones X and Y were not used in the training but their scoring results are clear: X is predicted to be normal, while Y is predicted to have a genomic aberration. After performing the analysis, the experimental team revealed to the analysis team that this prediction is correct.
Conclusion: The VAIDR classification method correctly predicted the genomic status.
Further and more detailed studies and data sets are necessary to show the robustness of the prediction across different cell lines and different genotypes.

“We have successfully supported many projects using iPSCs, both in drug discovery as well as cell therapy applications. If you have an application in mind, sign up for a free trial now!”
–Bruno Chilian, CSO at TRI